Lysozyme is an antibacterial enzyme with a molecular weight of 14.4kDa and is a member of the innate immune system. It can catalyze the 1-cleavage between N-acetylmuramic acid and N-acetylglucosamine residues in bacterial cell wall peptidoglycan. Hydrolyzes the 4-β-glycosidic bond, destroying the bacterial cell wall. Since the peptidoglycan layer of Gram-positive bacteria is much thicker than that of Gram-negative bacteria and is the main component of its cell wall, this enzyme mainly targets Gram-positive bacteria.
Lysozyme is contained in some human cell secretions, such as saliva, tears, and nasal discharge; lysozyme is also found in cytoplasmic granules and egg whites in mitochondria.
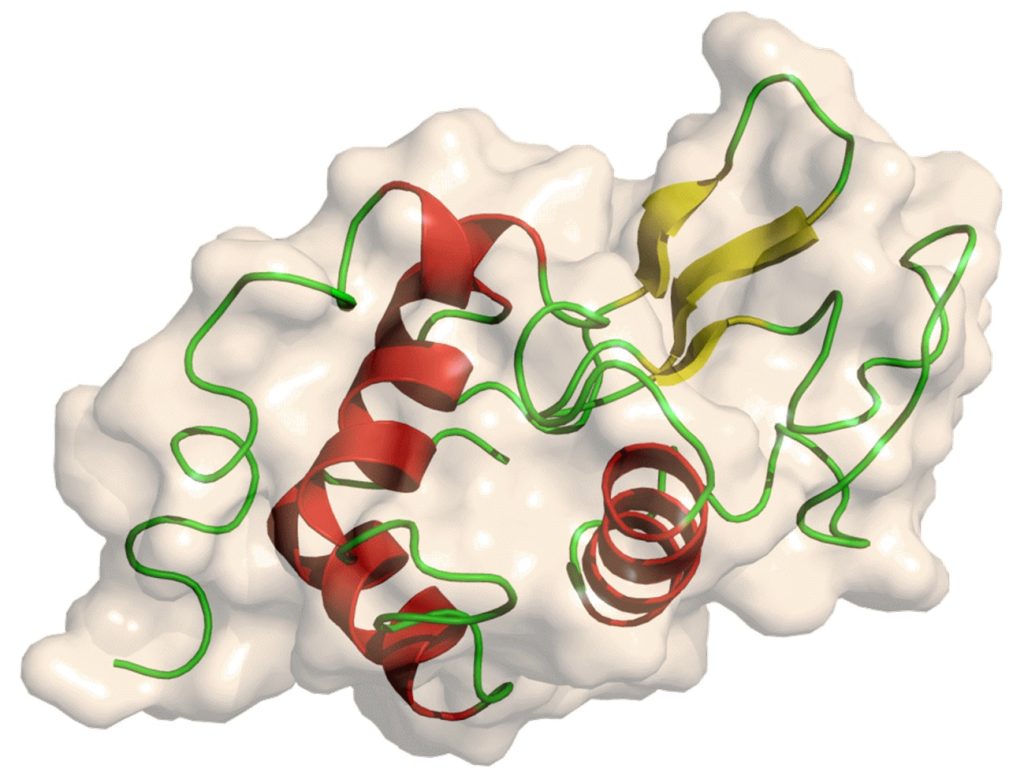
In 1909, Laschtschenko discovered that there were bactericidal substances in egg whites. In 1922, Alexander Fleming discovered and named lysozyme while studying the antibacterial effects of nasal mucus.
In 1965, David Philips used X-ray diffraction technology to study lysozyme crystals and solved the crystal structure with a resolution of 2 angstroms. This is the second protein structure to be obtained using X-ray diffraction technology and the first enzyme structure to be solved. David Philips proposed a mechanism of lysozyme catalysis based on the substructure, which has recently been revised.

Physiological Function
Lysozyme is part of the body’s immune system and kills Gram-positive bacteria. Lysozyme binds to bacterial surfaces, reducing negative charge and assisting in phagocytosis of bacteria. Lack of lysozyme in newborns can lead to bronchopulmonary dysplasia. Infants consuming formula that lacks lysozyme are three times more likely to develop diarrhea. A lack of lysozyme in tears can lead to conjunctivitis.
Certain cancer cells secrete excessive amounts of lysozyme, leading to high levels of lysozyme in the blood, leading to kidney failure and hypokalemia.
Catalytic Mechanism
Lysozyme attacks peptidoglycan (a component of bacterial cell walls, especially abundant in the cell walls of Gram-positive bacteria) and hydrolyzes the glycosidic bond connecting the fourth carbon atom of N-acetylmuramic acid and N-acetylglucosamine. The whole process is that lysozyme first binds to the peptidoglycan molecule through the “groove” between its two domains; then its substrate forms a transition state conformation in the enzyme. According to the Phillips mechanism, lysozyme binds to glucan. Lysozyme then twists the fourth sugar on the glucan into a half-chair conformation. In this twisted state (higher energy), glycosidic bonds can easily break.
The side chains of glutamic acid (Glu35) and aspartic acid (Asp52) at position 52 of the lysozyme protein sequence were found to be critical for lysozyme activity. Glu35 serves as a proton donor for glycosidic bonds and cleaves the C-O bond of the substrate; while Asp52 serves as a nucleophile and participates in the generation of glycosylase intermediates. Subsequently, the glycosylase intermediate reacts with water molecules and is hydrolyzed to produce the product, while the enzyme remains unchanged.
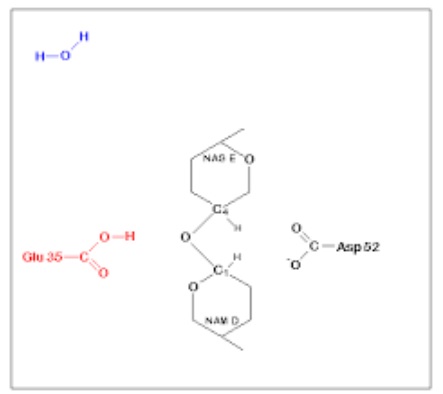
The mechanism of Lysozyme reaction during the hydrolysis of the beta (1-4) glycosidic
bond between N-acetylglucosamine sugar (NAG) and N-acetylmuramic acid sugar (NAM).
Related Diseases
In some forms of hereditary amyloidosis, it is found that patients contain a mutation in the gene encoding lysozyme, which causes lysozyme to accumulate in multiple tissues in the body.
Application
- Lysozyme is widely used in the laboratory for cell disruption of bacteria.
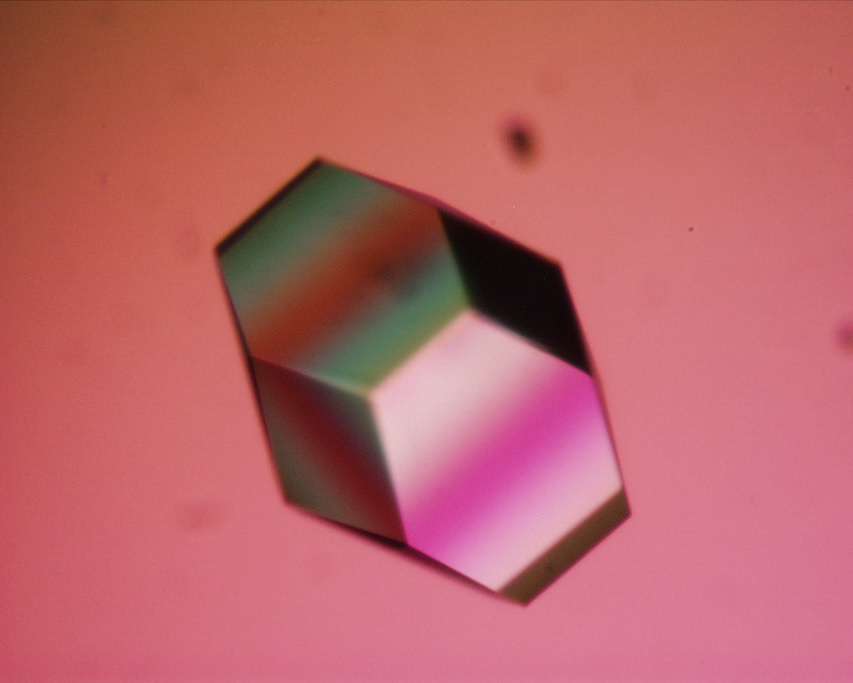
- Because lysozyme is easy to crystallize, it is often used in various crystallography-related studies.



